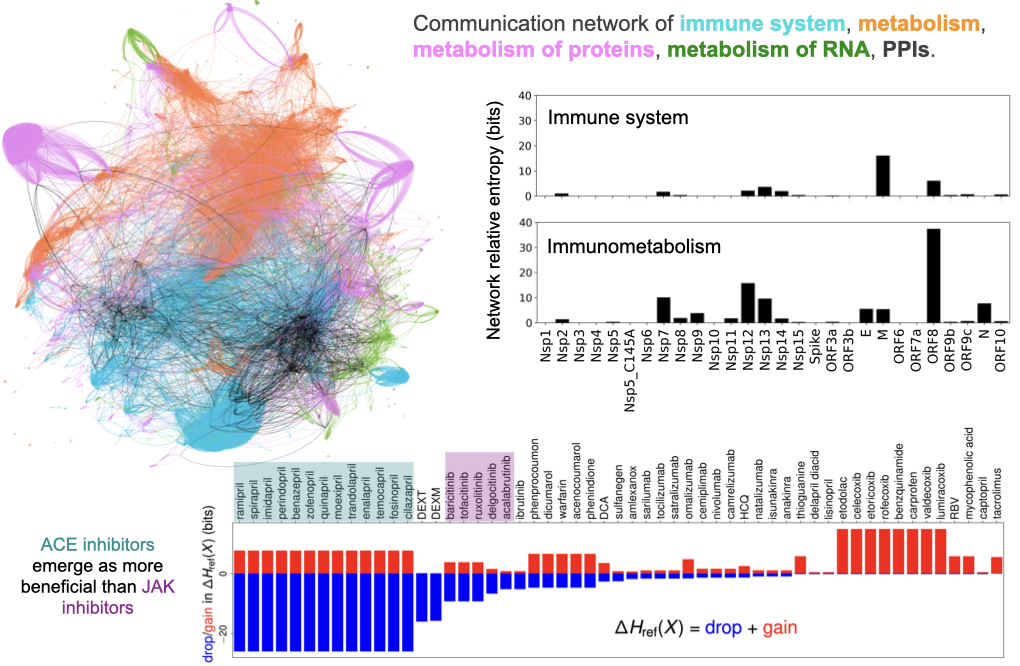
Communication network models for biological pathways
I developed Communication through Reactome and Interactome (CoRe), a computational framework to model biological pathways as information channels and compute the effective information reaching the proteins and metabolites through biochemical signaling. CoRe mines the Reactome pathways database for the reactions and the Human Reference Interactome database for protein-protein interactions. The image below shows the communication network model built using four top-level pathways from the Reactome database and the protein-protein interactions. The relative network entropy shows the information transferred by the SARS-CoV-2 proteins. We can use CoRe to evaluate the efficacy of candidate drugs to reduce the information transfer from the viral proteins.
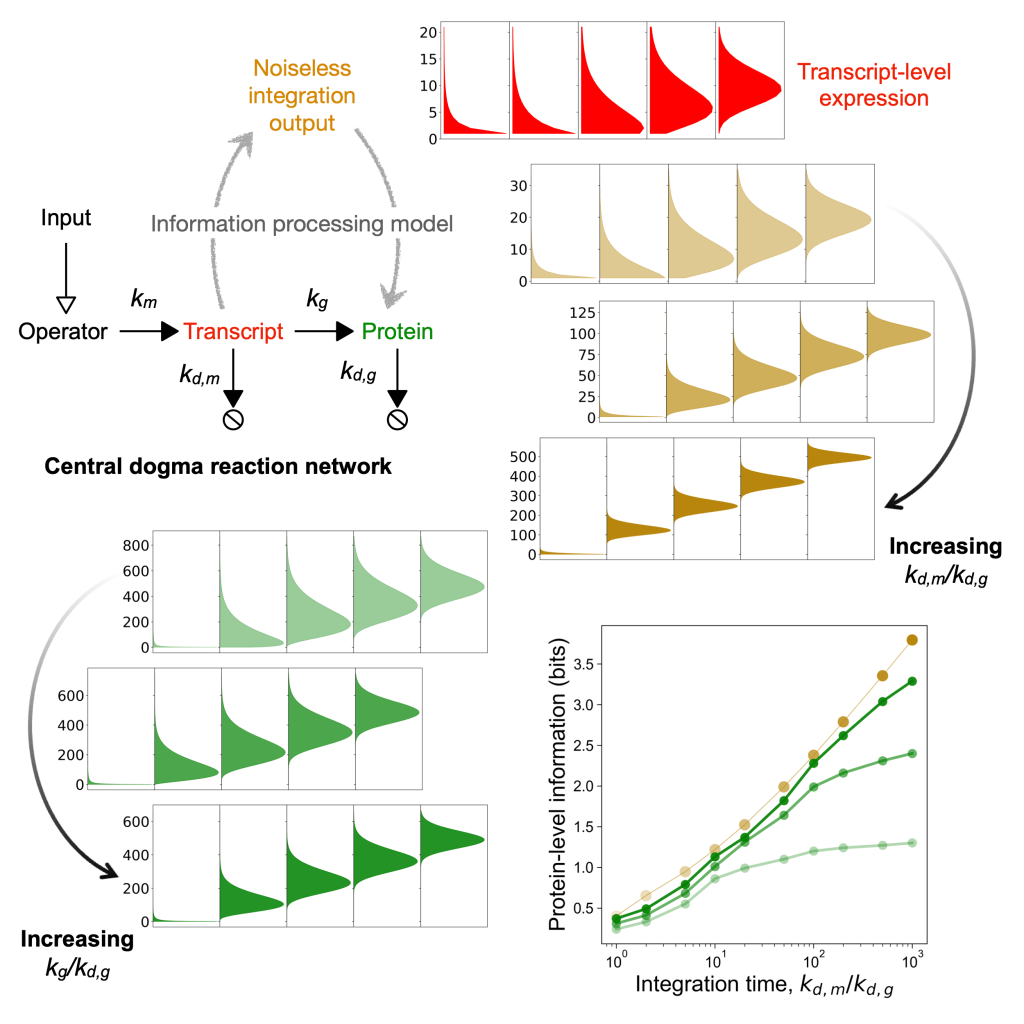
Nearly maximal information gain due to time integration in central dogma reactions
The central dogma reaction network gains information about an environmental input using the processes of transcription, transcript decay, translation, and protein decay. Both experimental and simulation data show that the information about an environmental input is higher in the protein expression level compared to the transcript expression level, which occurs due to time integration during translation. Using the central dogma rate constants data from four species, E. coli, M. musculus, S. cerevisiae, and H. sapiens, we showed that life achieves the maximum information gain possible due to time integration while keeping the information loss due to stochasticity in translation low.
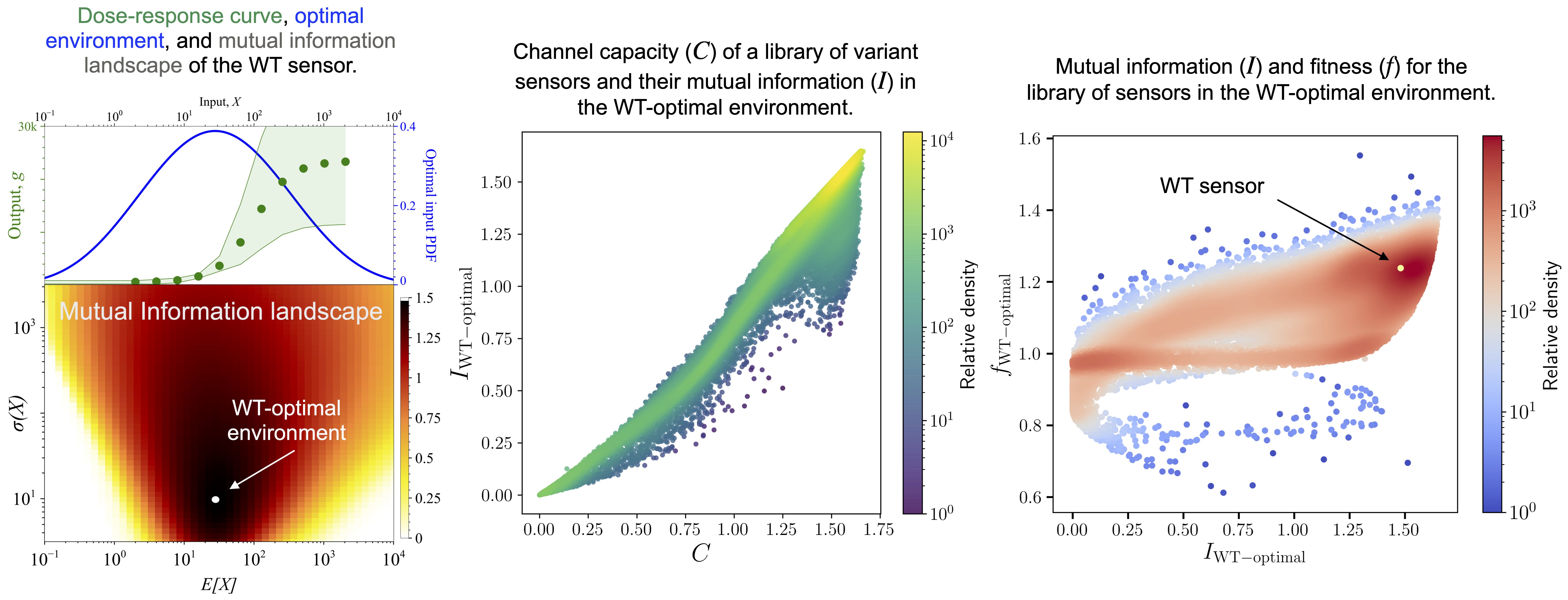
Information transfer and fitness in large library of sensor variants
Characterization of mutual information for a library of genetic sensor variants reveals the importance of information transfer on fitness in the wild-type (WT) optimal environment. The distribution of the variants in the mutual information and fitness space identifies variants that achieve optimal information transfer.
Information thermodynamics of large gene networks
The topology of regulatory interactions determines the phenotypic states that exist in these biological processes like differentiation, apoptosis, and homeostasis. The efficacy of communication in gene regulatory networks (GRNs) determine thermodynamic properties relevant for phenotypic transitions, like the inducible free energy change due to master regulators.

From 2013-2018 I worked on polymer networks: stochastic reaction-diffusion kinetics, topological properties, crosslink clustering, and collective modes in disordered networks.
Information transfer through biochemical reaction networks (BRNs)
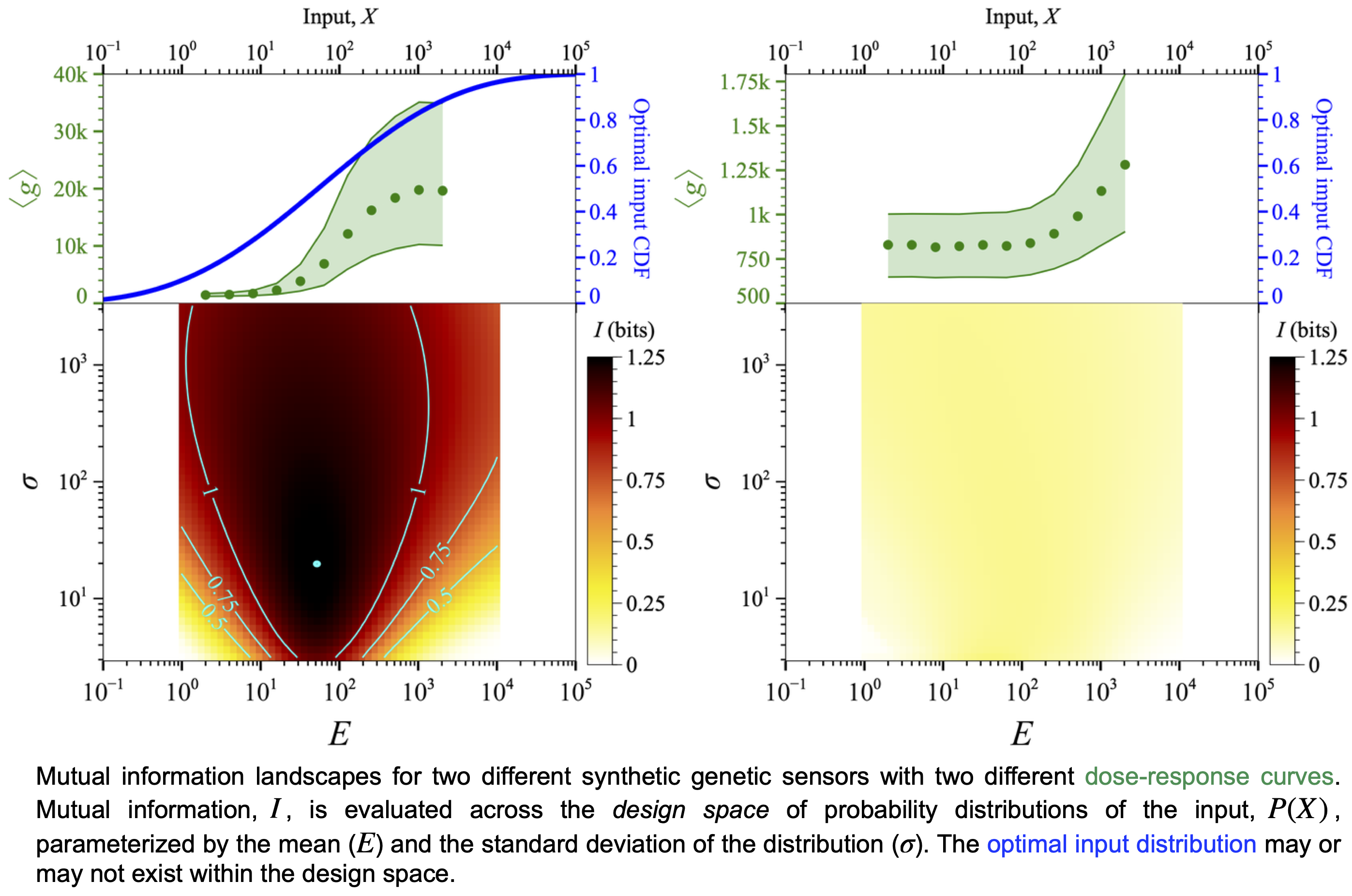
Transfer of information from the extracellular environment to the cellular response is essential for metabolic and regulatory functions. Sparse Estimation of Mutual Information Landscapes (SEMIL) evaluates the mutual information (I) between the input and the output of BRNs across the space of probability distributions of the input. Mutual information landscapes quantitatively illustrate the effect of feedback, stronger or weaker repression, etc. on the information transfer through BRNs.
Stochastic network growth during photopolymerization
Photopolymerization is the method of initiating a polymerization reaction using light energy, extensively used for fast 3D printing. During the reaction the monomer molecules connect together to form a polymer network.

The following movie demonstrates polymerized network growth using the stochastic reaction-diffusion model of the process. The green pixels are radicals, the red ones are network nodes, and the yellow pixels are crosslinks.
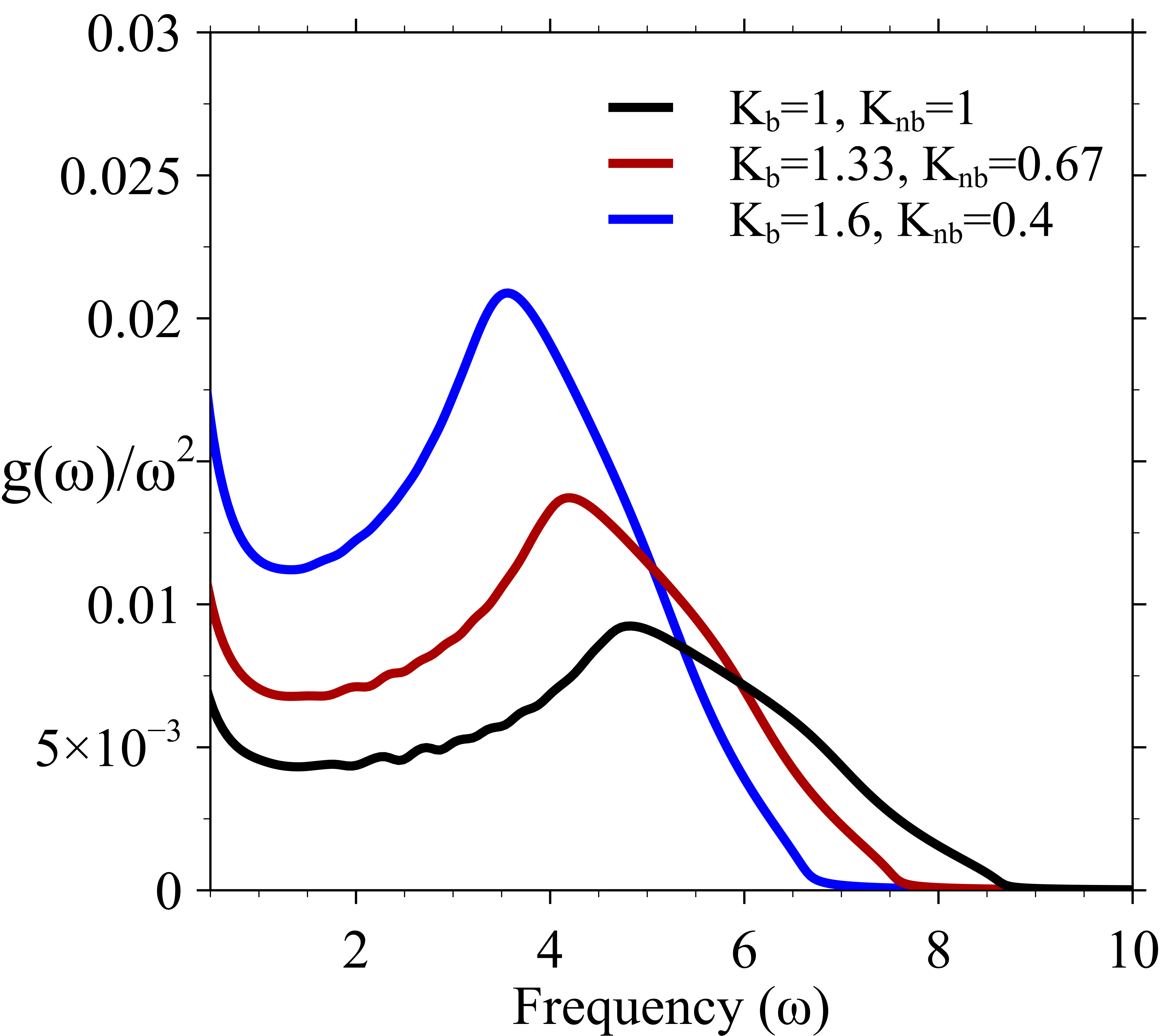
Phonon properties of disordered networks
Physical properties of polymers, like modulus and thermal conductivity, depend on the collective vibrational (or phonon) modes of the disordered network. Depending on the relative strengths of the bonded and non-bonded interactions, the same network structure can result in different probability distributions for force constants (K) for harmonic motion. The distribution in force constant is used to determine the phonon density of states.. The figure at left below shows the reduced density of states that are obtained for the three different distributions of force constants. The boson peak moves to lower frequencies and the relative strength of the bonded interaction increase with respect to non-bonded interaction.